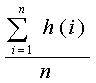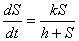 .
.
The three types of cells in the simulation,
astrocytes, microglia and neurons, all secrete
and absorb chemicals. Astrocytes and microglia
can move. It is possible to explore the effects
of cell division and death of microglia, though
these are omitted in the default setting.
In the current simulation, there are two types of absorption
models, Michaelis-Menten kinetics and receptor kinetics.
Michaelis-Menten Kinetics
Michaelis-Menten kinetics describe the rate of absorption as a
function of the concentration of chemical present, S. Two
parameters are necessary for determining the rate of absorption, the
maximum absorption rate (call it k) and the concentration
of chemicals at which the rate of absorption is half its maximum (call
it h). Then the change in concentration S can be
described by the following differential equation:
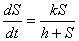 .
.
A slightly more detailed model for absorption kinetics tries to
describe the process through receptor kinetics. Let r be the
concentration of receptors for a given cell. Then by using the
kinetics of this receptor, specifically the rate at which the chemical
S binds to the receptor, kf, and the rate at which
it unbinds, kb, one can formulate a differential
equation through the laws of mass action to describe the binding
kinetics. If one were to assume that half of the receptors are active
(the proportion of active receptors actually varies), then the change
in concentration S can be described by the following differential
equation:
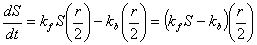
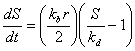 .
.
However, the latest addition of the simulation relaxes the
assumption that half of the receptors are bound. In this case,
external chemicals, So, can be absorbed into cells
to become internal, Si, by binding to receptors,
r, as shown in the following reaction:
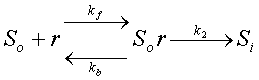 .
.The initial microglia count dictates the number of microglial agents initially in the simulation. The microglia concentration designates how many microglia are represented per agent. Thus, if the intial count is 40 and the concentration is 10, then 400 microglia are represented in the simulation. By using the concentration parameter one can change the number of microglia in the simulation without changing the computation power. The microglial agent is placed (or centered) within a grid space. The number of agents centered in any one grid space cannot exceed the maximum density parameter.
At initialization, a random pairing of (x,y) is generated.
If there is room (meaning that the maximum density parameter
will not be exceeded) to place a microglial agent in this grid space,
a microlgial agent will be placed in the grid space. If there is
no room, a new random pairing of (x,y) is generated and
tested. This process continues until a number of microglial agents
equal to the initial microglia count are in place.
Microglia absorb soluble amyloid according to the
Michaelis-Menten kinetics discussed above. The maximum
sAB microglia uptake rate is k and the half max
amyloid binding conc is h. Because this occurs on
the macro time scale, the amount of
soluble amyloid changed in the same grid space as the microglial
agent, call it DS, is
Microglia can also ingest amyloid fibers at a constant rate.
The programmer defines a fiber ingestion fraction, z
(currently, z=0.1), that is multiplied with the maximum
sAB microglia uptake rate, k to determine the constant
rate of ingestion. Because this occurs on the
macro time scale, the amount of fibrous amyloid, F, changed
in the same grid space as the microglial agent, call it DF, is
In addition to absorbing soluble amyloid, it is assumed that
microglia can neutralize the concentration of soluble amyloid that
it has stored. It does this at a constant rate defined by the
sol-AB neutralization rate parameter.
In this case, the change in concentration within the cell
is simply the product of the microglia concentration, the
neutralization rate and the macro time step. Of course, care
is taken so that the concentration of soluble amyloid in storage
is never negative.
Microglia secrete IL-1B. For details on how this is done, please
see IL-1B under
Chemicals.
The motility time delay parameter determines how many macro time steps a microglial agent must wait before moving. If there is no delay, then the speed of movement is simply the length of a grid space, DX, divided by the macro time step, DT. In general, if m is the number of steps between movement, then the corresponding speed is v = DX/(DT+1).
Each microglial agent has a counter which gets decremented every time step. When the counter reaches zero the cell attempts to move according to the rules specified below. Also at this time, the counter gets reset to the motility time delay value. When microglial cells are initialized, the counter takes any integer value between zero and the time delay, uniformly distributed. This allows for movement to be asynchronous, meaning that not all microglia will move at the exact same time.
Movement of microglia can be hampered by the presence of amyloid
fibers. If the concentration of fibers in the grid space of the
microglial cell is F and the concentration of fibers at
which a cell has a 50-50 chance of being stuck is called G
(this is the half-sticking fiber level parameter), then the
the probability, P, that a cell gets stuck is
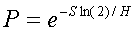 .
.
Microglia move chemotactically up the soluble amyloid gradient.
Thus, the first step in cell movement is to determine
which of the surrounding grid spaces has the greatest concentration
of soluble amyloid. The grid spaces sampled can be labeled as
0 1 2
3 4 5
6 7 8
where 4 is the current grid space of the microglial agent.
In the case where the cell is on a boundary, periodic boundary conditions
are assumed. After sampling the immediate neighborhood, the direction
of greatest concentration is noted. The cell will move in this
direction in a Monte Carlo fashion with the probability of
winning given by the chemotactic sensitivity parameter.
(Simply put, the cell will move in the direction of highest
concentration with a probability equal to the chemotactic sensitivity
parameter.) If the designated direction loses or all concentrations
are the same so that there are no designated directions, then a
direction 0-8 is chosen at random, uniformly distributed.
Having finally decided on a direction of movement, the cell must
check to see if room is available for movement. Room exists if no
astrocytes are in the grid space and the addition of another microglial
agent will not make the number of microglial agents in the grid space
exceed the maximum density parameter. If there is no room,
then the microglial agent will stay in its current grid space.
If cell death is turned on, then one of two factors will
determine whether a microglial agent will die. If the concentration
of absorbed soluble amyloid ever reaches the fatal sol-AB dosage
parameter, then the cell will die. A cell could also die based on
old age. We assume that the probability that a cell dies by age
T is based on a sigmoidal distribution,
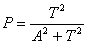
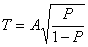 .
.If cell mitosis is turned on, then four criteria determine whether mitosis can occur. These factors are the length of cell cycle, half-sticking fiber level, poison effects on mitosis, and chance of mitosis. All four criteria must be satisfied in order for mitosis to occur.
The length of cell cycle determines the time at which mitosis can occur. Every microglial agent has a counter which has the cell's current age (in number of macro time steps since the cell was initialized). The length of cell cycle, call it C, is converted into number of macro time steps, M = C/DT. Whenever, the age counter is a multiple of M, mitosis can occur. Of course, this process can be changed, so that all microglial agents do not divide at the same time as follows: The length of the cell cycle can determine the mean of a normal distribution with the standard deviation another parameter to be added to the simulation. The timing for mitosis can then be handled much like cell death above, where each time a microglial agent is initiated, a time for cell mitosis can be randomly generated based on the normal distribution (or any other distribution). Cells can then keep track of a mitosis counter which gets decremented every macro time step. When the counter reaches zero, the other criteria are checked and the counter gets reset (stochastically based on the mitosis distribution like it was at initiation).
Because amyloid fibers are deemed hazardous to microglia, presence of fibers may prevent mitosis. In our current formulation, if the concentration of amyloid fibers in the same grid space as the microglial agent is greater than the half-sticking fiber level, then mitosis is prevented. If the concentration of amyloid fibers is less than the half-sticking fiber level, then the other criteria are checked.
The amount of soluble amyloid within the microglia (having been absorbed) also affects mitosis. The poison effects on mitosis parameter accounts for this. If the parameter equals one, then the cell is deemed poisoned if any soluble amyloid is stored inside the cell. If the parameter equals zero, then soluble amyloid does not act as a poison which prohibits mitosis. Values in between dictate a percentage of the capacity (the fatal sol-AB dosage parameter) at which the cell is poisoned. In this case, let E be the value of the poison effects parameter, S be the concentration of soluble amyloid within the cell, q be the microglia concentration and C be the capacity of soluble amyloid within a cell. The microglial agent is poisoned if S is greater than q(1-E)C in which case mitosis cannot occur. Otherwise, the other criteria are checked.
In our current formulation, all the other criteria are deterministic (the condtions are either met or not). To add a stochastic component to the model, the chance of mitosis parameter is checked in a Monte Carlo fashion. Thus, if all the other criteria have been met, a uniformly distributed random variable between 0 and 1 is generated. If this number is less than the chance of mitosis parameter, mitosis occurs. Otherwise, nothing happens until the next time mitosis can be checked.
If all the conditions for mitosis are met, then the program will
attempt to add another microglial agent to the simulation. It first
checks to make sure that there is enough memory (currently, the
simulation can hold 1000 microglial agents). A new microglial agent
is then initiated in the same grid space as the old one. The
concentration of soluble amyloid in storage in the old agent is
divided so that the old agent holds half and the new agent begins
with half. If the addition of the new microglial agent would make
the number of microglial agents exceed the maximum density
parameter, then the new agent simply replaces the old one.
The initial astrocyte count designates the number of astrocyte agents initially in the simulation. The astrocyte concentration designates how many astrocyte are represented per agent. Thus, if the intial count is 100 and the concentration is 10, then 1000 astrocytes are represented in the simulation. By using the concentration parameter, one can change the number of astrocytes in the simulation without changing the computation power. The astrocyte agent is placed (or centered) within a grid space. The number of agents centered in any one grid space cannot exceed the maximum density parameter.
At initialization, a random pairing of (x,y) is generated.
If there is room (meaning that the maximum density parameter
will not be exceeded, and that there are no fibers or microglial
agents already in the grid space) to place an astrocyte agent in
this grid space, one will be placed at (x,y).
If there is no room, a new random pairing of (x,y) is
generated and tested. This process continues until the number of
astrocyte agents placed equals the initial astrocyte count.
In the simulation, an astrocyte can be in one of four states:
As soon as the concentration of IL-1B in the same grid space
as the astrocyte agent exceeds the activation level,
the agent's state changes from inactive to receptive.
If the concentration ever falls underneath the activation level,
the state of the astrocyte will return to inactive as long
as it is not already blocking.
If the state of the astrocyte agent is not inactive, the agent
can absorb IL-1B. Absorption is based on the
receptor kinetics as described above. The
Il-1B receptor binding rate is kf. It
is used in conjunction with the IL-1B receptor equilibrium
constant to determine kb. The IL-1B
receptors per astrocyte is converted by the program
(via a programmer defined conversion constant) into a
concentration, R. Finally, the IL-1B receptor conversion
rate defines k2. Using these parameters,
the program can calculate k and h as defined in
the receptor kinetics (case 2). Thus, if
S is the concentration of IL-1B present, then the astrocyte
will attempt to remove an amount, DS, on the
macro time scale
(time increment = DT) such that
Astrocytes secrete IL-6 and TNF based on the fraction of bound
IL-1B receptors. As described in the section of
neuron health, the fraction of bound IL-1B receptors is given
by
In order for astrocyte movement to occur, the state of the astrocyte must either be receptive or motile. In addition to having the proper state, the amount of IL-1B within the astrocyte agent must exceed the product of the IL-1B threshold for motility and the astrocyte concentration.
The maximum speed parameter, v, is used to determine the number of macro time steps between movement, m. By letting DX be the length of a grid space and DT be the macro time increment, then m is the rounded integer value of (DX/v)/DT. Every astrocyte agent has an internal counter which keeps track of the number of macro time steps until movement can occur. Initially, the counter is set randomly between 0 and m (via a uniform distribution on the integer values). Every macro time step, the counter is decremented until it reaches zero. At this point, movement can occur and the counter is reset to m.
Astrocytes try to move towards deposits of amyloid fiber. The
astrocyte agent may move in one of several directions based on
the grid. These directions are labeled 0-8 as follows
0 1 2
3 4 5
6 7 8
where 4 determines the current position of the cell.
Different rules apply to receptive and motile
astrocyte agents.
Receptive astrocytes sample the following grid spaces
for amyloid fiber:
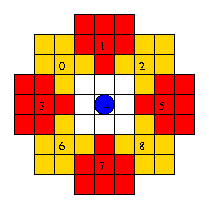
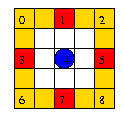
Even if movement should take place, it can only occur provided that there is room for the astrocyte agent in the new grid space. Room in the new grid space is based on three criteria: (1) no microglia can be present, (2) no amyloid fibers can be present and (3) the number of astrocyte agents cannot exceed the maximum density parameter.
If no fiber concentration is detected, then movement will not
occur. If the state of the astrocyte is motile when this
happens, then the state of the astrocyte will revert back to
receptive.
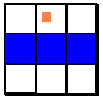
Any astrocytes that are not inactive may become what
we call blockers. In order for an astrocyte to become
a blocker, some amyloid fiber must be in a neighboring grid
space (these are the white spaces shown in the pictures for
movement above). If fibers are present,
then the astrocytes change the diffusivities in the immediate
grid spaces normal to the direction at which the fiber was
detected. The following pictures give examples of which grid
spaces are affected given that the astrocyte agent is located
at the center and the fiber is in the same grid space as the
orange square.
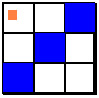
Unlike microglia and astrocytes, neurons cannot move. The current formulation assumes that there is a neuron in every grid space. Earlier formulations assumed that multiple grid spaces lead to what was called a neuron block which is simply a population of neurons over some rectangular block of grid spaces. The computer code for this approach has not been deleted, but neither has it been updated.
The major functions of neurons in the simulation are to absorb IL-1B, IL-6 and TNF, secrete soluble amyloid, and change health.
At initialization, the neuron located in the grid space in
the center of the environment
begins with a concentration of IL-1B equal to the average
of the maximum IL-1B absorbed parameter and the source
triggering level parameter. In this
manner, the center grid space will always be a source of soluble
protein. This allows the feedback loop to begin and result
in a meaningful simulation.
Absorption of IL-1B, IL-6 and TNF are all handled by the
receptor kinetics described above.
Therefore, for each chemical that is to be absorbed, there
are corresponding user controlled parameters associated with
them: the chemical receptor binding rate is kf;
by multiplying it with the chemical receptor equilibrium
constant, one obtains kb; the chemical
receptor conversion rate is k2; and the
chemical receptors per astrocyte is converted by the program
(via a programmer defined conversion constant) into a
concentration, R. Using these parameters,
the program can calculate k and h as defined in
the receptor kinetics (case 2).
If S is the concentration of chemical (be it IL-1B, IL-6
or TNF) in the same grid space as the neuron, then the
change in concentration, DS can be calculated on the
macro time scale
with time increment, DT, based on numerically solving
the differential equation. Using a simple Euler method, we
find that
Special consideration is given when the chemical reaches the neuron's capacity to store it. The neuron will never absorb more chemical than it can hold. The limit for IL-1B is given by the maximum IL-1B absorbed parameter. The limit for IL-6 is given by the fatal IL-6 concentration parameter. The limit for TNF is given by the maximum TNF absorbed parameter. In the case where DS exceeds the room available to the chemical, DS is decreased to take the remaining room.
In the unlikely case that DS exceeds S (the
concentration of chemical that is present in the grid space), DS
is changed so that it equals S. This is just a check
to make sure that no negative concentrations of chemicals
result from the absorption process. This makes sense both
biologically and mathematically.
Neurons act as sources for soluble amyloid protein based on the amount of IL-1B they have absorbed. In order for a neuron to become a source of soluble amyloid, the concentration of IL-1B that it has absorbed must exceed the source triggering level parameter. At the time at which this occurs, a final test is done to see if it actually becomes a source based on the maximum proportion of neuron sources parameter, m. This "test" is done in a Monte Carlo fashion where a uniformly distributed random variable between 0 and 1 is generated. If its value is less than m, the "test" is passed and the neuron becomes a source of soluble amyloid. Because, IL-1B is not reduced within the neuron, this "test" is only done once and will determine whether the neuron will be a source for the duration of the simulation. Of course, neuron death causes any sources of soluble amyloid to die with the neuron.
For details on the amounts of soluble amyloid secreted,
see Amyloid Protein
under Chemicals.
Neuron health varies as a rate depending on the proportion of bound receptors and local amyloid concentrations. Health varies between 0 and 1. A value of 0 for health indicates death. A value of 1 indicates that the neuron is in perfect health.
Because neuron health varies as a rate, we describe changes
in neuron health based on a differential equation. The
first assumption we make is that as long as neurons do
not decay past a certain point, the threshold of no
recovery, they can recover via the logistic equation,
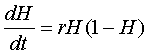
We then assume that the cytokines, IL-1B, IL-6 and TNF can
change health depending on the fraction of bound receptors.
The fraction of bound receptors can be determined from the
Michaelis-Menton kinetics. If A
the concentration of chemical in the same grid space as the
neuron, we can determine the fraction of bound receptors
for the chemical, BA, as
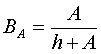
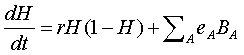
Finally we assume that the amyloid concentrations can
affect health as well. We allow both the local concentration
of soluble amyloid, S, and fibrous amyloid, F,
to influence neuron health. Again we incorporate an
effects on health parameter for soluble and fibrous
amyloid, eS and eF,
respectively. We make one final addition that scales the
concentrations, so that the full effects won't be felt unless
the concentration of S and F are at the
amyloid conc affecting health, Smax,
and fiber conc affecting health, Fmax,
respectively. Combining all of these effects, the final
equation for neuron health is
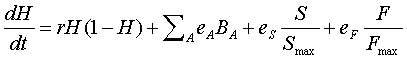
We calculate neuron health on the macro time scale with time increment, DT, based on numerically solving the above differential equation, using a simple Euler method. H can never exceed 1 so that if the calculation is greater than 1, then H is automatically set to 1. Likewise, H can never be less than zero (the neuron is dead when H=0) so that if the calculation is less than zero, then H is automatically set to zero.
Averaging the healths of
all the neurons gives one the overall neuron health.
Thus, if h(i) is the health of neuron i
and there are n neurons, then the overall neuron
health is simply